 
You can create a jupyter notebook (for this it’s good to open a new screen in your terminal) with the following command Then I add anaconda3 to my path by modifying the. I did this using the following commands (on my Putty terminal): wget -P /tmp īash /tmp/Anaconda3-2020.02-Linux-x86_64.sh Ok I will post the process I followed here for future users who want to set this up (if there are corrections/alternatives you may reply to this post).įirst I installed anaconda on my directory because I want to be able to install any package I want (also anaconda comes with ipython and jupyter). Perhaps it could be a useful workflow for you as well. Nevertheless, I find the setup quite useful until I debug code and parameters for ordinary job submission. That is probably also a reason why interactive jobs resources are limited to something like 47 cores. Note this setup is prone to errors of allocating resources longer than necessary as it requires to exit the job manually. For Julia, I can use the machinefile with julia -machine-file=$PBS_NODEFILE which will initiate workers and connect them with the master instance. Unfortunately, I can not understand why the first results on google with machinefile and python leads to MPI. When that is done, you can feed the machinefile for most systems. The next step is to make ssh passwordless between all two nodes for which I did use a neat bash line which I, unfortunately, can’t recall. You shall be able to ssh in any of the specified nodes. You can get the cores allocated to you in the machinefile which can be listed on hpc05 with cat $PBS_NODEFILE. To start using it, you can execute on the cluster qsub -I -l nodes=2 which will start the job and will put you in one of the nodes. But I am not sure how to set it up so any help by someone with experience would be greatly appreciated!Īn easy way to start getting used to with the cluster is an interactive mode. 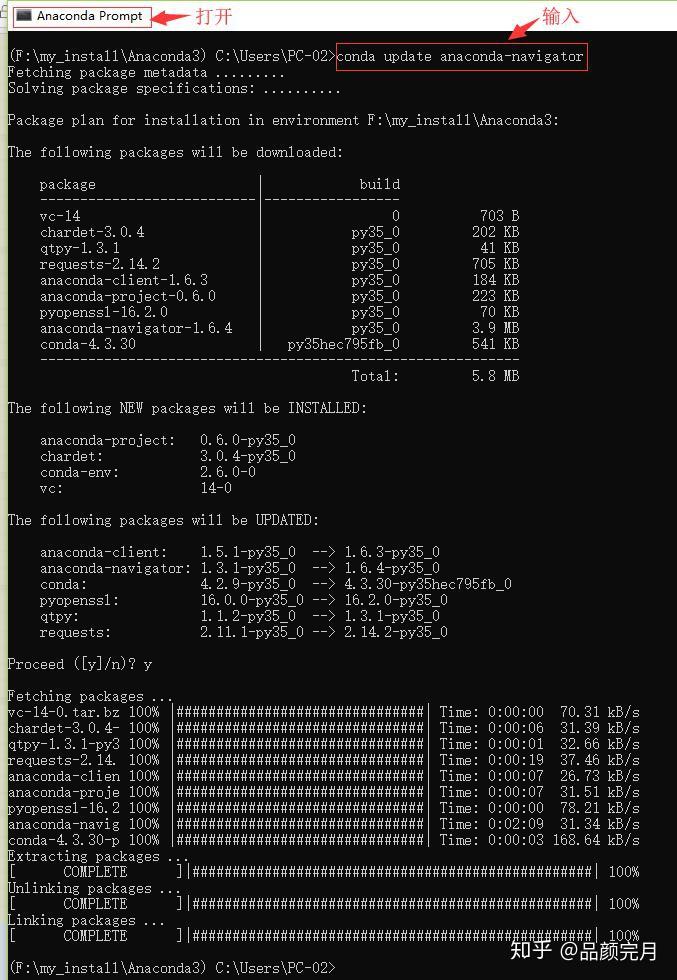
If running jupyter notebooks is much more complicated than running python scripts I could also go for that option. I would also like to compute P(t,x,y) while sweeping one of the parameters in the equation, which is what the cluster is really helpful for as far as I understand. 
So this is where the cluster could help speed things up. However, the quantity I am really interested in is the correlation function, which requires the knowledge of the conditional probability density P(t,x,y|t’,x’,y’), which basically means that I have to run my simulation above N by M times (for each initial condition P(t’,x’,y’)=1). (This can successfully run on my laptop, although I need to severely constrain my grid size due to memory issues). The result I get is the probability density P(t,x,y) for all t, x and y, which I can use to calculate for example. I discretise the equation in the spatial dimensions (N by M grid) so I have a system of N by M ordinary differential equations which are solved using a python ode solver. I numerically solve a 2D Fokker-Planck equation which is basically a partial differential equation containing a partial derivative with respect to time and partial derivatives with respect to x and y. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
